Unlocking the Human Proteome
Northwestern scientists are leading the charge to sequence proteins the way the Human Genome Project sequenced genes and DNA. A recent study in Science confirms they are on to something big.
by Gina Bazer

Neil Kelleher, PhD, has been weighing proteins since 1999, when, as a young professor, he opened his independent laboratory at the University of Illinois in Urbana-Champaign. His goal then, he says, was the same as his goal now: “to weigh every human protein.”
More specifically, he is aiming to measure the exact molecular composition, or unique proteoform, of 100 million proteins to understand and treat diseases ranging from cancer to neurogenerative and heart diseases. At last count, he’s classified about 200,000 unique proteoforms, leaving many millions more to go. This will require a similar, nearly 1,000-fold jump in sequencing technology as occurred in the Human Genome between 1992 and 2002, when the genome was first fully sequenced, he says.
Fortunately, Kelleher, who is now a professor of Medicine at Feinberg and the Walter and Mary E. Glass Professor of Molecular Biosciences and professor of Chemistry in the Weinberg College of Arts and Sciences, is no longer the only one counting. In November of 2021, approximately 400 scientists in a worldwide proteomics research consortium (topdownproteomics.org) launched an initiative known as the Human Proteoform Project. Kelleher serves as the founding president.

“People told me I was leaning in far over my skis when I started, but now momentum is growing steadily,” says Kelleher, who is also the director of the Chemistry of Life Processes Institute (CLP) and faculty director of Northwestern Proteomics, a center within CLP that helps clinicians define the proteoform signatures of health and disease. “There are technological leaps that need to be made, and there are many signs that show this could be the decade where it finally happens.”
Watch this space, both in the private and public sectors.
NEIL KELLEHER, PhD
“Today, there is a realization that unlocking protein-level biology is standing between us and a far healthier and more efficient future for interacting with our own proteomes,” Kelleher says. “After the genome project, it’s an obvious next step toward precision medicine, as proteins underlie all our health and every disease. Our bodies are driven by proteoform biology, so we are talking about the earlier and more precise detection of all human disease, better drug development, and improving healthy lifespans through regenerative medicine.”

While still in its early stages, Kelleher and his colleagues’ mission to map all proteins is hardly an abstraction. Recently, their efforts moved conceptually from bench to bedside. A recent study published in Science describes how Kelleher and his lab, partnering with Northwestern Medicine transplant hepatologist Josh Levitsky, MD, mapped 57,000 new proteoforms and discovered families of proteins in the body that could potentially predict which patients might reject a new liver transplant. The advancement, which Kelleher describes as microcosm of the Human Proteoform Project’s macro potential, marks the beginning of a new era for proteomics.
A Revolutionary Top-Down Analysis of Protein Patterns
Conventionally, scientists have tended to look at shifting patterns of proteins as if through goggles underwater, taking in just a fraction of available information about their unique structures through a “bottom-up” analysis that digests whole proteins into peptides and identifies what remains. The Kelleher lab does a “top-down” analysis using state-of-the-art mass spectrometry to identify proteoforms in cells and blood more precisely (and efficiently) by keeping them intact, rather than cutting them up into tiny pieces.
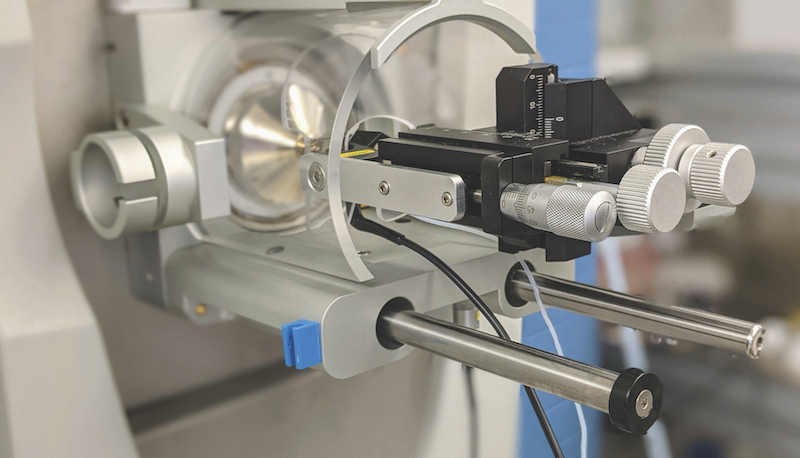
“Bottom-up leaves major knowledge gaps,” says Kelleher, which is why, more than 25 years ago, he led the charge to develop top-down proteomics. At the time, he used a machine the size of a VW bus containing a superconducting magnet, created by his mentor, the late Fred McLafferty, PhD, of Cornell University. “I would manually process each step, including the measurement of intact proteins with a five-ton instrument resembling an MRI machine, and it would take me a whole month to characterize a single proteoform,” Kelleher recalls.

Fast forward to the present. The Kelleher lab now uses a dozen benchtop instruments that weigh about 200 pounds each. The technology has come a long way from 2012, when Kelleher published his first case for mapping the entire human proteome in the Journal of the American Society of Mass Spectrometry. More modern instruments allow scientists in this growing field to measure a few thousand proteoforms in a week. Kelleher predicts scientists will begin mapping proteoforms 100 times more rapidly over the next five to 10 years, using a variety of new technologies now in their infancy.
“Watch this space, both in the private and public sectors,” he says.
For the Science paper, Kelleher’s top-down method allowed the scientists in his lab to take a magnifying glass to 21 human cell types and blood to create a map of protein families. They then held the map up in front of liver transplant recipients and found new indicators in immune cell proteins that changed with acute organ rejection. The result, the Blood Proteoform Atlas (BPA) — “just one specific-use case and a small pilot for the Human Proteoform Project,” according to Kelleher — advances the field by 10-fold over any previous study.
A Blood Test for Liver Transplant Rejection
Having team members across disciplines allows the project to conceptualize a clinical application. As Kelleher probes the scientific basis for phenomena in the cell, he works with Levitsky, the study’s co-corresponding author, to understand how these could be applied to a specific system. Levitsky, who is also a professor of Medicine, Surgery and Medical Education, originally connected with Kelleher through his leadership in the biomarkers space, in which measurable signs in the blood are used to predict health metrics in patients facing disorders — and in this instance, liver transplant rejection.
“It was really important for Neil that there was a biologically relevant example to contextualize how these proteoform panels can identify diseases non-invasively as markers,” Levitsky says. “And there’s also a need in my field to have mechanistic biomarkers that are more relevant to their immune biological pathways. This could be the start of a new era in cell-specific markers.”
Physicians must suppress the immune system with drug therapy and monitor liver transplant recipients for signs of rejection, often only detected after an episode has begun. Guesswork throughout this process could be eliminated with specific knowledge about what’s happening at the most granular level.
This could be the start of a new era in cell-specific markers.
JOSH LEVITSKY, MD
With the BPA as a reference map, the team took blood samples from participants in one of Levitsky’s biomarker collection studies. They examined which proteoforms seemed to activate in response to the transplant and identified those that changed compared to patients without rejection.
Next, the Levitsky and Kelleher team developed a panel of 24 proteoforms from the initial study and looked at them in transplant recipient samples from across the country. They found the same proteoforms lit up as in the first trial.
“The promise here is to be able to use this panel moving forward to be able to identify patients who have no signs of rejection versus those who have very early evidence of rejection,” Levitsky says. “If we can pick up on this several weeks before rejection actually happens, we might be able to modify immunosuppression.”
Levitsky continues to examine how proteoforms change in transplant recipients over time to develop additional biomarkers that may inform how he treats patients down the line. Kelleher says as the number of cell types in the atlas grows, so, too, will potential ways to use it. In addition to broadening understandings of human biology, the BPA could have similar applications across immune disorders.
Endless Possibilities
Each human gene can have from just a few to sometimes over 100 unique proteoforms. And with 20,300 individual genes in the human body, there are millions of proteoforms created by modification or splicing. Kelleher says with a reasonably complete roadmap of each gene’s family of proteins, discoveries about disease, aging, and new therapeutics will accelerate significantly.
“We are trying to communicate this potential to the world,” Kelleher says. “Feinberg and Northwestern are really at the forefront of this thinking, and I feel fortunate that we have been able to come as far as we have. The arc of history is on our side.”
Listen to a podcast with Kelleher here.
Additional reporting by Win Reynolds
Proteoforms in Cardiovascular Disease
The story of cardiovascular disease risk is often boiled down to the traditional risk factors: cholesterol, blood pressure, insulin resistance, body weight, tobacco use, diet, physical activity, and genetics. But they don’t explain all the risk, according to John T. Wilkins, MD, MS, associate professor of Medicine in the Division of Cardiology and of Preventive Medicine in the Division of Epidemiology. He says another factor could be modifications to apolipoproteins, a category of proteins found in the blood that mediate cholesterol metabolism.
Using participants from the Coronary Artery Risk Development in Young Adults (CARDIA) study, Wilkins and Kelleher measured proteoforms of apolipoprotein A1 (apoA1) in 150 participants, comparing specific ratios of proteoforms to cardiovascular risk factors such as waist circumference (a marker of abdominal body fat) and HDL cholesterol. Publishing their findings in the Journal of the American Heart Association, the investigators reported a positive association between HDL — good cholesterol — and several proteoform variants of apoA1.
“These findings present opportunities to enhance the understanding of metabolic health and potentially develop better biomarkers,” Wilkins says. – Will Doss